# You'll start seeing this cell in most lectures.
# It exists to hide all of the import statements and other setup
# code we need in lecture notebooks.
from dsc80_utils import *
Agenda¶
numpyarrays.- From
babypandastopandas.- Deep dive into DataFrames.
- Accessing subsets of rows and columns in DataFrames.
.locand.iloc.- Querying (i.e. filtering).
- Adding and modifying columns.
pandasandnumpy.
We can't cover every single detail! The pandas documentation will be your friend.
Question 🤔 (Answer using `lec02-dog`)
dogs = pd.read_csv('data/dogs43.csv')
dogs.head(2)
What does this code do?
whoa = np.random.choice([True, False], size=len(dogs))
(dogs[whoa]
.groupby('size')
.max()
.get('longevity')
)
numpy arrays¶
numpy overview¶
numpystands for "numerical Python". It is a commonly-used Python module that enables fast computation involving arrays and matrices.numpy's main object is the array. Innumpy, arrays are:- Homogenous – all values are of the same type.
- (Potentially) multi-dimensional.
- Computation in
numpyis fast because:- Much of it is implemented in C.
numpyarrays are stored more efficiently in memory than, say, Python lists.
- This site provides a good overview of
numpyarrays.
We used numpy in DSC 10 to work with sequences of data:
arr = np.arange(10)
arr
# The shape (10,) means that the array only has a single dimension,
# of size 10.
arr.shape
2 ** arr
Arrays come equipped with several handy methods; some examples are below, but you can read about them all here.
(2 ** arr).sum()
(2 ** arr).mean()
(2 ** arr).max()
(2 ** arr).argmax()
⚠️ The dangers of for-loops¶
for-loops are slow when processing large datasets. You will rarely writefor-loops in DSC 80 (except for Lab 1 and Project 1), and may be penalized on assignments for using them when unnecessary!- One of the biggest benefits of
numpyis that it supports vectorized operations.- If
aandbare two arrays of the same length, thena + bis a new array of the same length containing the element-wise sum ofaandb.
- If
- To illustrate how much faster
numpyarithmetic is than using afor-loop, let's compute the squares of the numbers between 0 and 1,000,000:- Using a
for-loop. - Using vectorized arithmetic, through
numpy.
- Using a
%%timeit
squares = []
for i in range(1_000_000):
squares.append(i * i)
In vanilla Python, this takes about 0.04 seconds per loop.
%%timeit
squares = np.arange(1_000_000) ** 2
In numpy, this only takes about 0.001 seconds per loop, more than 40x faster! Note that under the hood, numpy is also using a for-loop, but it's a for-loop implemented in C, which is much faster than Python.
Multi-dimensional arrays¶
While we didn't see these very often in DSC 10, multi-dimensional lists/arrays may have since come up in DSC 20, 30, or 40A (especially in the context of linear algebra).
We'll spend a bit of time talking about 2D (and 3D) arrays here, since in some ways, they behave similarly to DataFrames.
Below, we create a 2D array from scratch.
nums = np.array([
[5, 1, 9, 7],
[9, 8, 2, 3],
[2, 5, 0, 4]
])
nums
# nums has 3 rows and 4 columns.
nums.shape
We can also create 2D arrays by reshaping other arrays.
# Here, we're asking to reshape np.arange(1, 7)
# so that it has 2 rows and 3 columns.
a = np.arange(1, 7).reshape((2, 3))
a
Operations along axes¶
In 2D arrays (and DataFrames), axis 0 refers to the rows (up and down) and axis 1 refers to the columns (left and right).
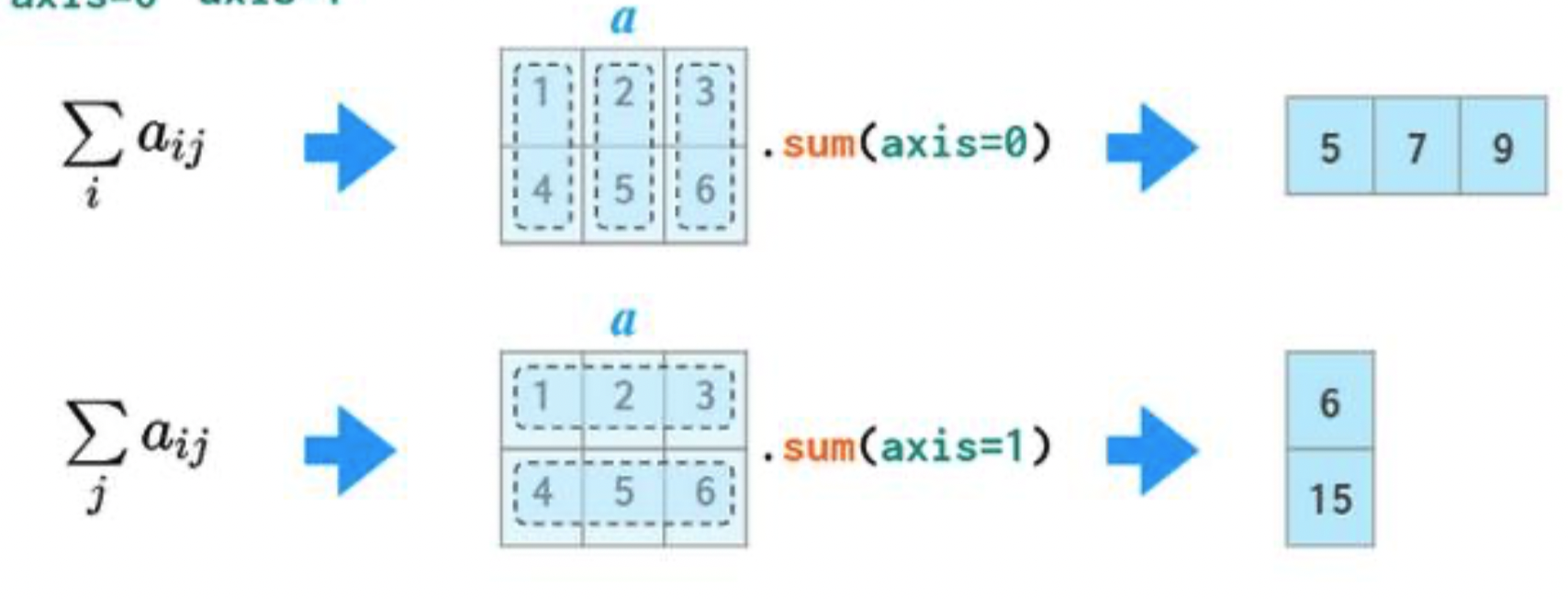
a
If we specify axis=0, a.sum will "compress" along axis 0.
a.sum(axis=0)
If we specify axis=1, a.sum will "compress" along axis 1.
a.sum(axis=1)
Selecting rows and columns from 2D arrays¶
You can use [square brackets] to slice rows and columns out of an array, using the same slicing conventions you saw in DSC 20.
a
# Accesses row 0 and all columns.
a[0, :]
# Same as the above.
a[0]
# Accesses all rows and column 1.
a[:, 1]
# Accesses row 0 and columns 1 and onwards.
a[0, 1:]
Question 🤔 (Answer at `lec02-grid`)
Try and predict the value of grid[-1, 1:].sum() without running the code below.
See if ChatGPT can do it.
s = (5, 3)
grid = np.ones(s) * 2 * np.arange(1, 16).reshape(s)
# grid[-1, 1:].sum()
Example: Image processing¶
numpy arrays are homogenous and potentially multi-dimensional.
It turns out that images can be represented as 3D numpy arrays. The color of each pixel can be described with three numbers under the RGB model – a red value, green value, and blue value. Each of these can vary from 0 to 1.
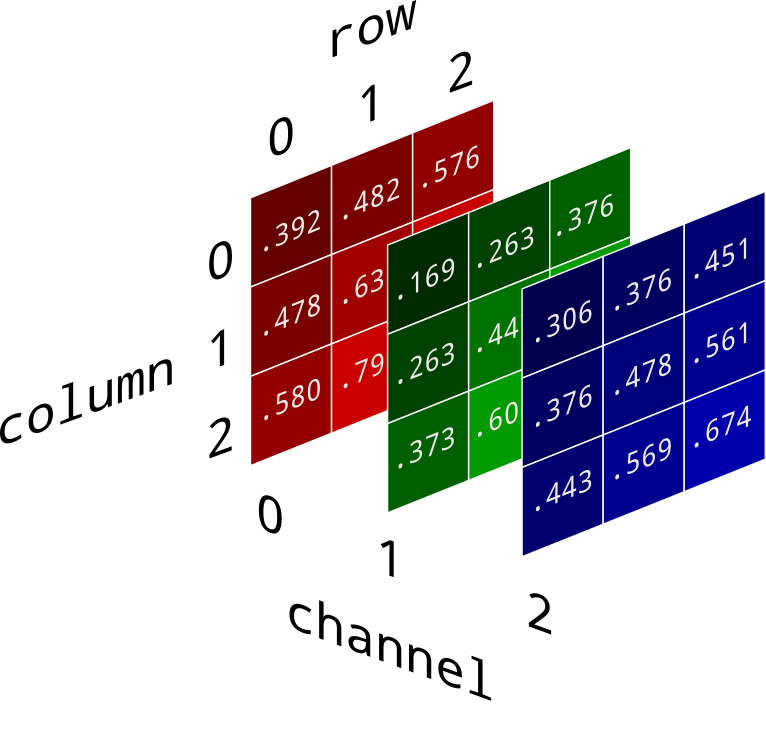 (image source)
(image source)from PIL import Image
img_path = Path('imgs') / 'bentley.jpg'
img = np.asarray(Image.open(img_path)) / 255
img
img.shape
plt.imshow(img)
plt.axis('off');
Applying a greyscale filter¶
One way to convert an image to greyscale is to average its red, green, and blue values.
mean_2d = img.mean(axis=2)
mean_2d
This is just a single red channel!
plt.imshow(mean_2d)
plt.axis('off');
We need to repeat mean_2d three times along axis 2, to use the same values for the red, green, and blue channels. np.repeat will help us here.
# np.newaxis is an alias for None.
# It helps us introduce an additional axis.
np.arange(5)[:, np.newaxis]
np.repeat(np.arange(5)[:, np.newaxis], 3, axis=1)
mean_3d = np.repeat(mean_2d[:, :, np.newaxis], 3, axis=2)
plt.imshow(mean_3d)
plt.axis('off');
Applying a sepia filter¶
Let's sepia-fy Junior!
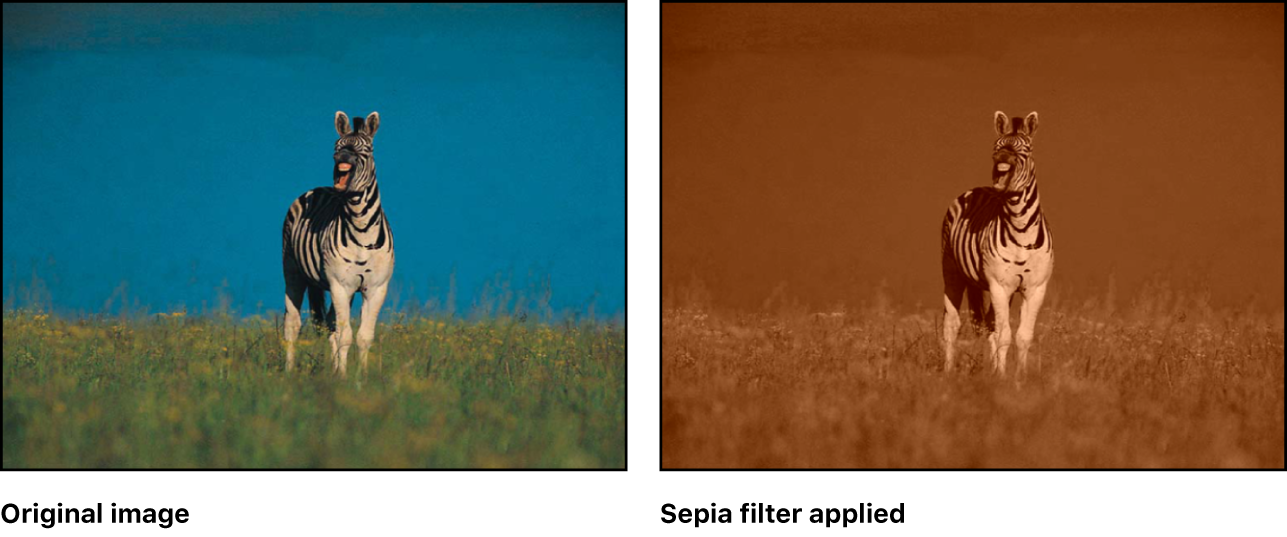 (Image credits)
(Image credits)
From here, we can apply this conversion to each pixel.
$$\begin{align*} R_{\text{sepia}} &= 0.393R + 0.769G + 0.189B \\ G_{\text{sepia}} &= 0.349R + 0.686G + 0.168B \\ B_{\text{sepia}} &= 0.272R + 0.534G + 0.131B\end{align*}$$
sepia_filter = np.array([
[0.393, 0.769, 0.189],
[0.349, 0.686, 0.168],
[0.272, 0.534, 0.131]
])
# Multiplies each pixel by the sepia_filter matrix.
# Then, clips each RGB value to be between 0 and 1.
filtered = (img @ sepia_filter.T).clip(0, 1)
filtered
plt.imshow(filtered)
plt.axis('off');
Key takeaway: avoid for-loops whenever possible!¶
You can do a lot without for-loops, both in numpy and in pandas.
From babypandas to pandas 🐼¶
babypandas¶
In DSC 10, you used babypandas, which was a subset of pandas designed to be friendly for beginners.

pandas¶
You're not a beginner anymore – you've taken DSC 20, 30, and 40A. You're ready for the real deal.

Fortunately, everything you learned in babypandas will carry over!
pandas¶

pandasis the Python library for tabular data manipulation.- Before
pandaswas developed, the standard data science workflow involved using multiple languages (Python, R, Java) in a single project. - Wes McKinney, the original developer of
pandas, wanted a library which would allow everything to be done in Python.- Python is faster to develop in than Java, and is more general-purpose than R.
pandas data structures¶
There are three key data structures at the core of pandas:
- DataFrame: 2 dimensional tables.
- Series: 1 dimensional array-like object, typically representing a column or row.
- Index: sequence of column or row labels.
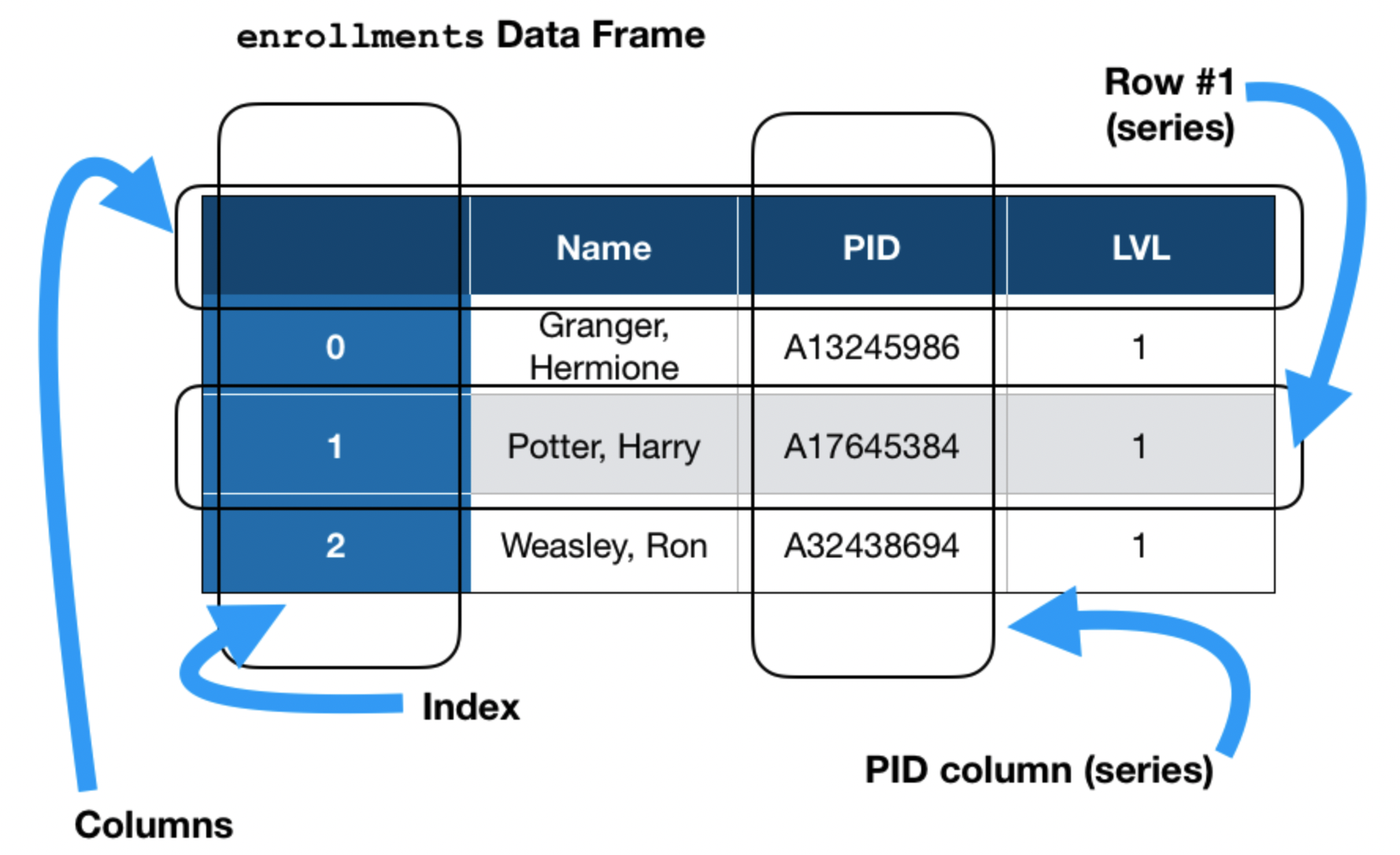 A DataFrame you'll see in Lab 1.
A DataFrame you'll see in Lab 1.
Importing pandas and related libraries¶
pandas is almost always imported in conjunction with numpy.
import pandas as pd
import numpy as np
# You'll see the Path(...) / subpath syntax a lot.
# It creates the correct path to your file,
# whether you're using Windows, macOS, or Linux.
dog_path = Path('data') / 'dogs43.csv'
dogs = pd.read_csv(dog_path)
dogs
Review: head, tail, shape, index, get, and sort_values¶
To extract the first or last few rows of a DataFrame, use the head or tail methods.
dogs.head(3)
dogs.tail(2)
The shape attribute returns the DataFrame's number of rows and columns.
dogs.shape
# The default index of a DataFrame is 0, 1, 2, 3, ...
dogs.index
We know that we can use .get() to select out a column or multiple columns...
dogs.get('breed')
dogs.get(['breed', 'kind', 'longevity'])
Most people don't use .get in practice; we'll see the more common technique in a few slides.
And lastly, remember that to sort by a column, use the sort_values method. Like most DataFrame and Series methods, sort_values returns a new DataFrame, and doesn't modify the original.
# Note that the index is no longer 0, 1, 2, ...!
dogs.sort_values('height', ascending=False)
# This sorts by 'height',
# then breaks ties by 'longevity'.
# Note the difference in the last three rows between
# this DataFrame and the one above.
dogs.sort_values(['height', 'longevity'],
ascending=False)
Note that dogs is not the DataFrame above. To save our changes, we'd need to say something like dogs = dogs.sort_values....
dogs
Setting the index¶
Think of each row's index as its unique identifier or name. Often, we like to set the index of a DataFrame to a unique identifier if we have one available. We can do so with the set_index method.
dogs.set_index('breed')
# The above cell didn't involve an assignment statement,
# so dogs was unchanged.
dogs
# By reassigning dogs, our changes will persist.
dogs = dogs.set_index('breed')
dogs
# There used to be 7 columns, but now there are only 6!
dogs.shape
Question 🤔 (Answer at `lec02-index`)
Ask ChatGPT:
- To explain what happens if you have duplicate values in a column and use
set_index()on it.
💡 Pro-Tip: Displaying more rows/columns¶
Sometimes, you just want pandas to display a lot of rows and columns. You can use this helper function to do that:
from IPython.display import display
def display_df(df, rows=pd.options.display.max_rows, cols=pd.options.display.max_columns):
"""Displays n rows and cols from df."""
with pd.option_context("display.max_rows", rows,
"display.max_columns", cols):
display(df)
display_df(dogs.sort_values('weight', ascending=False),
rows=43)
Selecting columns¶
Selecting columns in babypandas 👶🐼¶
- In
babypandas, you selected columns using the.getmethod. .getalso works inpandas, but it is not idiomatic – people don't usually use it.
dogs
dogs.get('size')
# This doesn't error, but sometimes we'd like it to.
dogs.get('size oops!')
Selecting columns with []¶
- The standard way to select a column in
pandasis by using the[]operator. - Specifying a column name returns the column as a Series.
- Specifying a list of column names returns a DataFrame.
dogs
# Returns a Series.
dogs['kind']
# Returns a DataFrame.
dogs[['kind', 'size']]
# 🤔
dogs[['kind']]
# Breeds are stored in the index, which is not a column!
# dogs['breed']
dogs.index
dogs
# What are the unique kinds of dogs?
dogs['kind'].unique()
# How many unique kinds of dogs are there?
dogs['kind'].nunique()
# What's the distribution of kinds?
dogs['kind'].value_counts()
# What's the mean of the 'longevity' column?
dogs['longevity'].mean()
# Tell me more about the 'weight' column.
dogs['weight'].describe()
# Sort the 'lifetime_cost' column. Note that here we're using sort_values on a Series, not a DataFrame!
dogs['lifetime_cost'].sort_values()
# Gives us the index of the largest value, not the largest value itself.
dogs['lifetime_cost'].idxmax()
Selecting subsets of rows (and columns)¶
Use loc to slice rows and columns using labels¶
You saw slicing in DSC 20.
loc works similarly to slicing 2D arrays, but it uses row labels and column labels, not positions.
dogs
# The first argument is the row label.
# ↓
dogs.loc['Pug', 'longevity']
# ↑
# The second argument is the column label.
As an aside, loc is not a method – it's an indexer.
type(dogs.loc)
type(dogs.sort_values)
.loc is flexible 🧘¶
You can provide a sequence (list, array, Series) as either argument to .loc.
dogs
dogs.loc[['Cocker Spaniel', 'Labrador Retriever'], 'size']
dogs.loc[['Cocker Spaniel', 'Labrador Retriever'], ['kind', 'size', 'height']]
# Note that the 'weight' column is included!
dogs.loc[['Cocker Spaniel', 'Labrador Retriever'], 'lifetime_cost': 'weight']
dogs.loc[['Cocker Spaniel', 'Labrador Retriever'], :]
# Shortcut for the line above.
dogs.loc[['Cocker Spaniel', 'Labrador Retriever']]
Review: Querying¶
- As we saw in DSC 10, querying is the act of selecting rows in a DataFrame that satisfy certain condition(s).
- Comparisons with arrays (or Series) result in Boolean arrays (or Series).
- We can use comparisons along with the
locoperator to filter a DataFrame.
dogs
dogs.loc[dogs['weight'] < 10]
dogs.loc[dogs.index.str.contains('Retriever')]
# Because querying is so common, there's a shortcut:
dogs[dogs.index.str.contains('Retriever')]
# Empty DataFrame – not an error!
dogs.loc[dogs['kind'] == 'beaver']
Note that because we set the index to 'breed' earlier, we can select rows based on dog breeds without having to query.
dogs
# Series!
dogs.loc['Maltese']
If 'breed' was instead a column, then we'd need to query to access information about a particular breed.
dogs_reset = dogs.reset_index()
dogs_reset
# DataFrame!
dogs_reset[dogs_reset['breed'] == 'Maltese']
Querying with multiple conditions¶
Remember, you need parentheses around each condition. Also, you must use the bitwise operators & and | instead of the standard and and or keywords. pandas makes weird decisions sometimes!
dogs
dogs[(dogs['weight'] < 20) & (dogs['kind'] == 'terrier')]
💡 Pro-Tip: Using .query¶
.query is a convenient way to query, since you don't need parentheses and you can use the and and or keywords.
dogs
dogs.query('weight < 20 and kind == "terrier"')
dogs.query('kind in ["sporting", "terrier"] and lifetime_cost < 20000')
Question 🤔 (Answer at `lec02-query`)
Ask ChatGPT:
- To explain when you would use
.query()instead of.loc[]or the other way around.
Don't forget iloc!¶
ilocstands for "integer location."ilocis likeloc, but it selects rows and columns based off of integer positions only, just like with 2D arrays.
dogs
dogs.iloc[1:15, :-2]
iloc is often most useful when we sort first. For instance, to find the weight of the longest-living dog breed in the dataset:
dogs.sort_values('longevity', ascending=False)['weight'].iloc[0]
# Finding the breed itself involves sorting, but not iloc.
dogs.sort_values('longevity', ascending=False).index[0]
More practice¶
Consider the DataFrame below.
jack = pd.DataFrame({1: ['fee', 'fi'],
'1': ['fo', 'fum']})
jack
For each of the following pieces of code, predict what the output will be. Then, uncomment the line of code and see for yourself. We may not be able to cover these all in class; if so, make sure to try them on your own. Here's a Pandas Tutor link to visualize these!
# jack[1]
# jack[[1]]
# jack['1']
# jack[[1, 1]]
# jack.loc[1]
# jack.loc[jack[1] == 'fo']
# jack[1, ['1', 1]]
# jack.loc[1,1]
Questions? 🤔
What questions do you have?
We will probably have to end lecture here.
Adding and modifying columns¶
Adding and modifying columns, using a copy¶
- To add a new column to a DataFrame, use the
assignmethod.- To change the values in a column, add a new column with the same name as the existing column.
- Like most
pandasmethods,assignreturns a new DataFrame.- Pro ✅: This doesn't inadvertently change any existing variables.
- Con ❌: It is not very space efficient, as it creates a new copy each time it is called.
dogs.assign(cost_per_year=dogs['lifetime_cost'] / dogs['longevity'])
dogs
💡 Pro-Tip: Method chaining¶
Chain methods together instead of writing long, hard-to-read lines.
# Finds the rows corresponding to the five cheapest to own breeds on a per-year basis.
(dogs
.assign(cost_per_year=dogs['lifetime_cost'] / dogs['longevity'])
.sort_values('cost_per_year')
.iloc[:5]
)
💡 Pro-Tip: assign for column names with special characters¶
You can also use assign when the desired column name has spaces (and other special characters) by unpacking a dictionary:
dogs.assign(**{'cost per year 💵': dogs['lifetime_cost'] / dogs['longevity']})
Adding and modifying columns, in-place¶
- You can assign a new column to a DataFrame in-place using
[].- This works like dictionary assignment.
- This modifies the underlying DataFrame, unlike
assign, which returns a new DataFrame.
- This is the more "common" way of adding/modifying columns.
- ⚠️ Warning: Exercise caution when using this approach, since this approach changes the values of existing variables.
# By default, .copy() returns a deep copy of the object it is called on,
# meaning that if you change the copy the original remains unmodified.
dogs_copy = dogs.copy()
dogs_copy.head(2)
dogs_copy['cost_per_year'] = dogs_copy['lifetime_cost'] / dogs_copy['longevity']
dogs_copy
Note that we never reassigned dogs_copy in the cell above – that is, we never wrote dogs_copy = ... – though it was still modified.
Mutability¶
DataFrames, like lists, arrays, and dictionaries, are mutable. As you learned in DSC 20, this means that they can be modified after being created. (For instance, the list .append method mutates in-place.)
Not only does this explain the behavior on the previous slide, but it also explains the following:
dogs_copy
def cost_in_thousands():
dogs_copy['lifetime_cost'] = dogs_copy['lifetime_cost'] / 1000
# What happens when we run this twice?
cost_in_thousands()
dogs_copy
⚠️ Avoid mutation when possible¶
Note that dogs_copy was modified, even though we didn't reassign it! These unintended consequences can influence the behavior of test cases on labs and projects, among other things!
To avoid this, it's a good idea to avoid mutation when possible. If you must use mutation, include df = df.copy() as the first line in functions that take DataFrames as input.
Also, some methods let you use the inplace=True argument to mutate the original. Don't use this argument, since future pandas releases plan to remove it.
pandas and numpy¶
pandas is built upon numpy!¶
- A Series in
pandasis anumpyarray with an index. - A DataFrame is like a dictionary of columns, each of which is a
numpyarray. - Many operations in
pandasare fast because they usenumpy's implementations, which are written in fast languages like C. - If you need access the array underlying a DataFrame or Series, use the
to_numpymethod.
dogs['lifetime_cost']
dogs['lifetime_cost'].to_numpy()
pandas data types¶
- Each Series (column) has a
numpydata type, which refers to the type of the values stored within. Access it using thedtypesattribute. - A column's data type determines which operations can be applied to it.
pandastries to guess the correct data types for a given DataFrame, and is often wrong.- This can lead to incorrect calculations and poor memory/time performance.
- As a result, you will often need to explicitly convert between data types.
dogs
dogs.dtypes
pandas data types¶
Notice that Python str types are object types in numpy and pandas.
| Pandas dtype | Python type | NumPy type | SQL type | Usage |
|---|---|---|---|---|
| int64 | int | int_, int8,...,int64, uint8,...,uint64 | INT, BIGINT | Integer numbers |
| float64 | float | float_, float16, float32, float64 | FLOAT | Floating point numbers |
| bool | bool | bool_ | BOOL | True/False values |
| datetime64 or Timestamp | datetime.datetime | datetime64 | DATETIME | Date and time values |
| timedelta64 or Timedelta | datetime.timedelta | timedelta64 | NA | Differences between two datetimes |
| category | NA | NA | ENUM | Finite list of text values |
| object | str | string, unicode | NA | Text |
| object | NA | object | NA | Mixed types |
This article details how pandas stores different data types under the hood.
This article explains how numpy/pandas int64 operations differ from vanilla int operations.
Type conversion¶
You can change the data type of a Series using the .astype Series method.
For example, we can change the data type of the 'lifetime_cost' column in dogs to be uint32:
dogs
# Gives the types as well as the space taken up by the DataFrame.
dogs.info()
dogs['lifetime_cost'] = dogs['lifetime_cost'].astype('uint32')
Now, the DataFrame takes up less space! This may be insignificant in our DataFrame, but makes a difference when working with larger datasets.
dogs.info()
💡 Pro-Tip: Setting dtypes in read_csv¶
Usually, we prefer to set the correct dtypes in read_csv, since it can help pandas load in files more quickly:
dog_path
dogs = pd.read_csv(dog_path, dtype={'lifetime_cost': 'uint32'})
dogs
dogs.dtypes
Axes¶
- The rows and columns of a DataFrame are both stored as Series.
- The axis specifies the direction of a slice of a DataFrame.
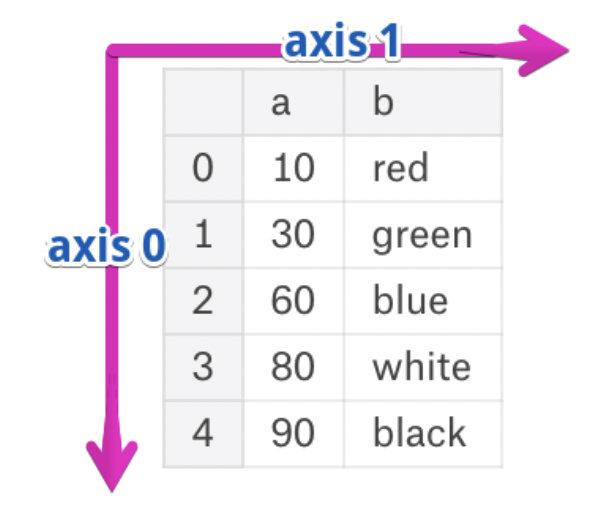
- Axis 0 refers to the index (rows).
- Axis 1 refers to the columns.
- These are the same axes definitions that 2D
numpyarrays have!
DataFrame methods with axis¶
- Many Series methods work on DataFrames.
- In such cases, the DataFrame method usually applies the Series method to every row or column.
- Many of these methods accept an
axisargument; the default is usuallyaxis=0.
dogs
# Max element in each column.
dogs.max()
# Max element in each row – throws an error since there are different types in each row.
# dogs.max(axis=1)
# The number of unique values in each column.
dogs.nunique()
# describe doesn't accept an axis argument; it works on every numeric column in the DataFrame it is called on.
dogs.describe()
Exercise
Pick a dog breed that you personally like or know the name of. Then:- Try to find a few other dog breeds that are similar in weight to yours in
all_dogs. - Which similar breeds have the lowest and highest
'lifetime_cost'?'intelligence_rank'? - Are there any similar breeds that you haven't heard of before?
For fun, look up these dog breeds on the AKC website to see what they look like!
all_dogs = pd.read_csv(Path('data') / 'all_dogs.csv')
all_dogs
# Your code goes here.
Summary, next time¶
Summary¶
pandasis the library for tabular data manipulation in Python.- There are three key data structures in
pandas: DataFrame, Series, and Index. - Refer to the lecture notebook and the
pandasdocumentation for tips. pandasrelies heavily onnumpy. An understanding of how data types work in both will allow you to write more efficient and bug-free code.- Series and DataFrames share many methods (refer to the
pandasdocumentation for more details). - Most
pandasmethods return copies of Series/DataFrames. Be careful when using techniques that modify values in-place. - Next time:
groupbyand data granularity.